People
Isaure Chauvot de Beauchêne
Principal investigator
Loria

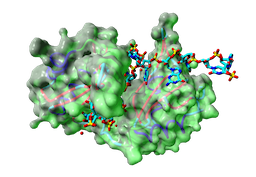
Project: Collect, integrate and analyse data on RNA recognition motifs (RRMs).
This project focuses on two main axes:
1. The creation of a complete and comprehensive database of available RRM information from the many available RRM data
covering a broad range of behaviours (with initial help of the other PhD students in the project). This includes
their sequence, structure, dynamics, RNA specificity and other data (binding affinity, biological function…).
This database will be regularly extended with internal and external data as it becomes available, will be released
at the end of the project, and is key to the development of computational approaches in the RNAct project.
You will analyse these different RRM data, and will liaise with the other PhD students to enrich the data with
results from in silico methodologies.
2. The computation of protein-RNA binding energies by molecular dynamics simulations of RRM-RNA models obtained by
your fellow- PhD students. For example, he/she will investigate why the Drosophilia Sex-lethal (Sxl) protein, and
its putative human homologue HuR, bind more specific Py-tracts than the U2AF65 protein and cannot accommodate cytosine.
This project involves a three-month secondment at VUB (Brussel, Belgium) to learn about sequence-based methods for protein design and analysis.
Isaure Chauvot de Beauchêne
Principal investigator
Loria

H2020-MSCA-ITN-2018 813239 (Jan. 1, 19-Dec. 31, 23)
RNAct - Enabling proteins with RNA recognition motifs for synthetic biology and bio-analytics